Student
Department of Chemical and Biochemical Engineering, University of Notre Dame.
Portada » Developing a pipeline for calcium image analysis

Student
Calcium is an essential element that helps orchestrate morphological processes, such as cell division and growth, during tissue development in many organisms At the cellular level, it acts as a “second messenger”, relaying biologically relevant signals from the environment and neighboring cells. The development of fluorescent molecules that make it possible to visualize local calcium concentrations under the microscope has been crucial to study calcium behavior in these processes and a great step forward for biology and biotechnology. This has allowed, for instance, to observe physiological changes occurring in a tissue over time.
Recent technical advances in fluorescence microscopy have enabled an even more detailed study of calcium dynamics, providing valuable insights into cellular responses to various stimuli and changes in their environment. The large amount of data generated from calcium microscopy imaging has also driven the development of technologies designed to extract, analyze, and interpret large volumes of information and its biological significance.
As part of The Bridge program, undergraduate student Gabriel Miranda, under the guidance of Professor Jeremiah Zartman from the University of Notre Dame, developed a pipeline for the analysis of calcium images using various computational tools to extract and quantify data related to the behavior of cells and tissues, using the fruit fly as a model organism, in a precise and highly automated manner.
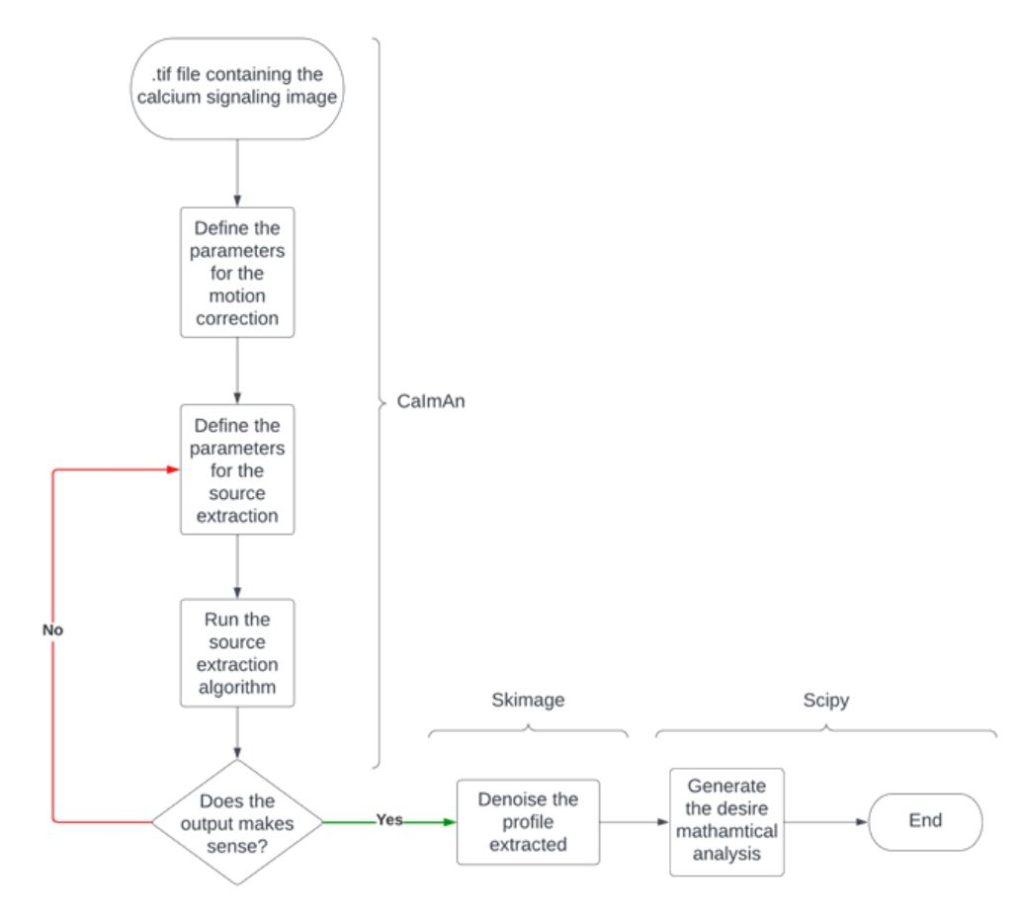
Workflow of the calcium imaging pipeline
The first step in the analysis consisted of applying motion correction to fluorescent calcium images using a series of algorithms. Next, they stored corrected images in a memory map, which involved source extraction to detect regions of interest. Finally, they carried out mathematical analyses to extract specific characteristics of calcium signals observed during the experiment, such as total number, peak height and width, and oscillatory frequency.
After completing the overall design, the researchers applied this pipeline to analyze experiments probing the development of fruit fly (Drosophila melanogaster) larvae conducted in Professor Zartman’s laboratory. In this experimental model, they study the function of Piezo, a membrane protein from the family of ion channels, which responds to mechanical stress by allowing calcium ions to flow into or out of cells. This channel translates mechanical forces into electrical signals, playing a crucial role in processes associated with the development and formation of different organs. Using Yoda1, a chemical compound that artificially activate Piezo channels, it was possible to study in more detail their role in the wing disc development and its related calcium dynamics. With this approach, they could demonstrate that calcium signals transmitted through Piezo are necessary to precisely regulate the size of this organ during the development of fly larvae, providing a better understanding of the role of this channel during embryonic development.
Thanks to the research conducted by Gabriel and Professor Zartman’s team, it is now possible to analyze calcium images and extract quantitative data automatically. This development facilitates the visualization, interpretation, and communication of results obtained from experiments studying calcium dynamics in cells and tissues. In this particular case, it provided insights into the development of fly wing tissue by experimentally altering the function of the Piezo channel. In addition, the code developed for this image analysis pipeline will allow researchers exploring calcium signals in various fields to perform detailed data analyses without requiring extensive programming or computer science knowledge.
Student
Department of Chemical and Biochemical Engineering, University of Notre Dame.

Student